
Description
uORF4u is a bioinformatics tool for conserved upstream ORF annotation.
Programming language: Python3
OS: MacOS, Linux
Python dependencies: biopython, configs, argparse, pandas, statistics, logomaker, matplotlib, reportlab.
Python version: >= 3.7
OS-level dependencies: mafft (v7.490 is included in the package)
License: CC0
Version: 0.9.6.1 (June 2025)
Web version is available at the Atkinson Lab Server Portal: server.atkinson-lab.com/uorf4u
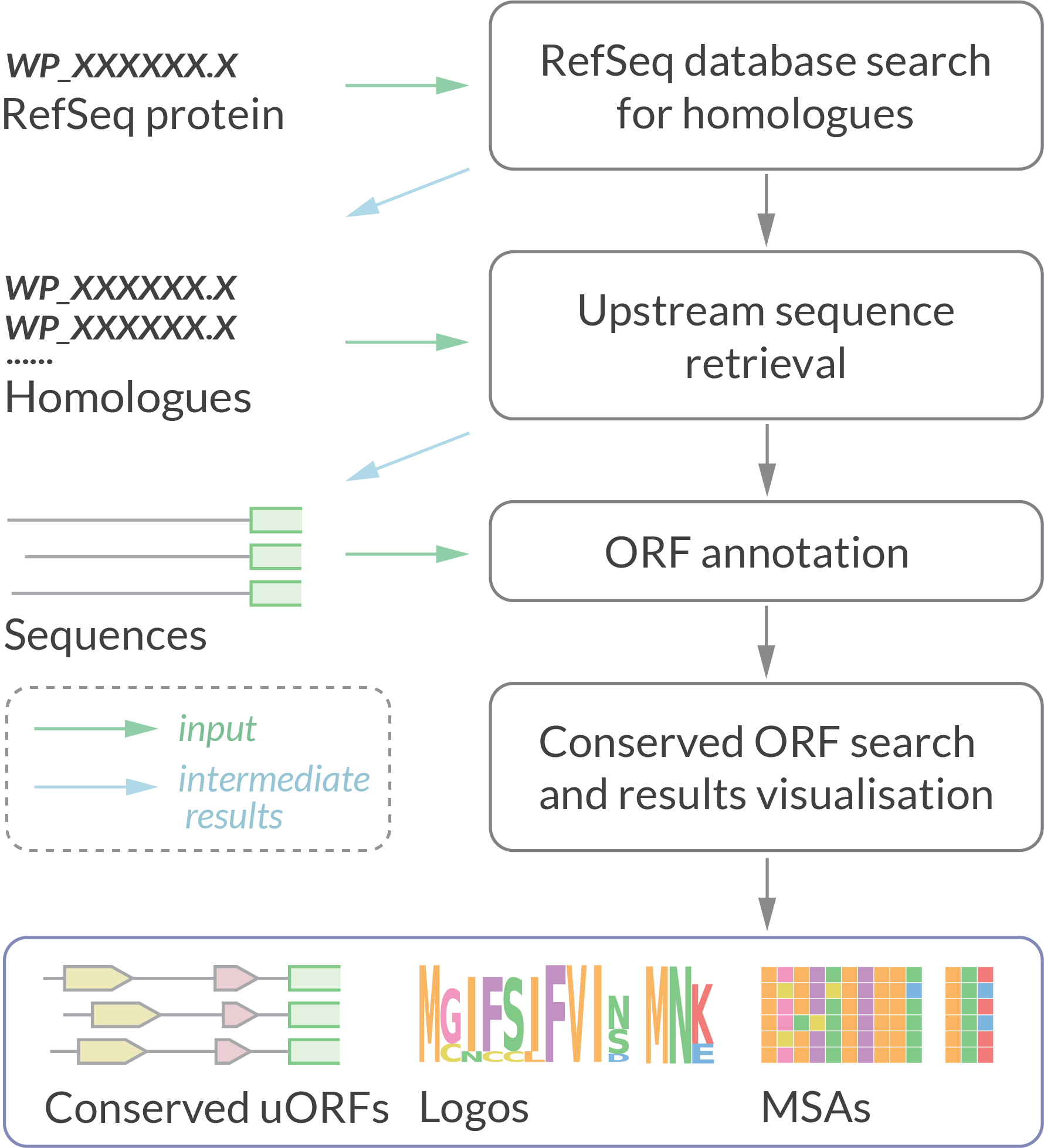
Data analysis pipeline
Installation
- The most stable release of uorf4u can be installed directly from pypi:
python3 -m pip install uorf4u
- The development version is available at github :
git clone https://github.com/GCA-VH-lab/uorf4u.git
cd uorf4u
python3 -m pip install --upgrade pip
python3 -m pip install wheel
python3 setup.py sdist bdist_wheel
python3 -m pip install -e .
! If you're a linux user, run uorf4u --linux post-install command once to update paths in the premade config files that set by default for MacOS users.
Reference
If you find uorf4u useful, please cite:
Artyom. A. Egorov, Gemma C. Atkinson, uORF4u: a tool for annotation of conserved upstream open reading frames, Bioinformatics, Volume 39, Issue 5, May 2023, btad323; doi: 10.1093/bioinformatics/btad323
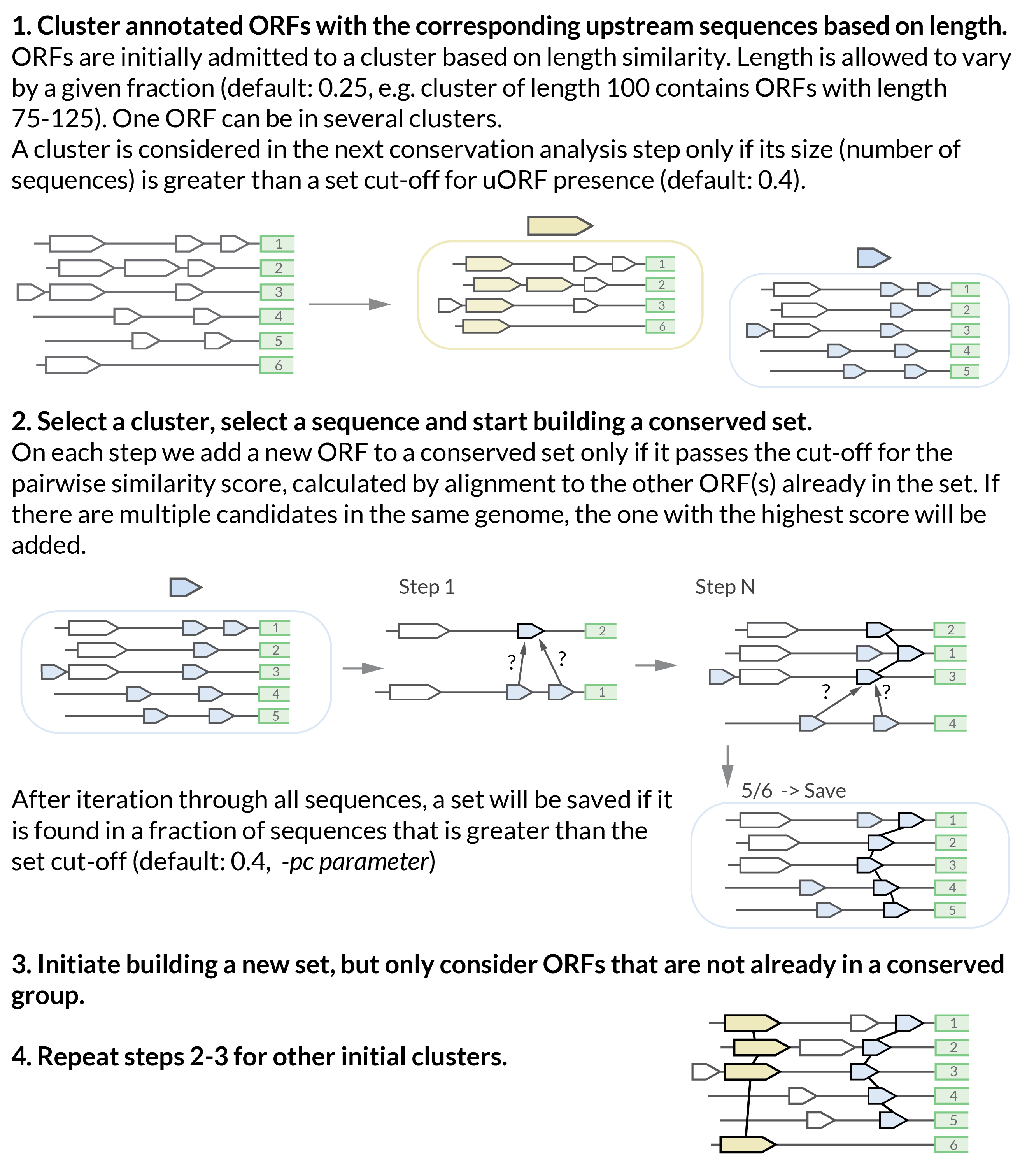
Conservation analysis algorithm
Contact
Please contact us by e-mail artemdotegorovATmeddotludotse or use Issues to report any technical problems.
You can also use Discussions section for sharing your ideas or feature requests!
Authors
uORF4u is developed by Artyom Egorov at the Atkinson Lab, Department of Experimental Medical Science, Lund University, Sweden 🇸🇪.
We are open for suggestions to extend and improve uorf4u functionality. Please don't hesitate to share your ideas or feature requests.